use http://www.philender.com/courses/data/hsbdemo, clear
regress write read
Source | SS df MS Number of obs = 200
-------------+------------------------------ F( 1, 198) = 109.52
Model | 6367.42127 1 6367.42127 Prob > F = 0.0000
Residual | 11511.4537 198 58.1386552 R-squared = 0.3561
-------------+------------------------------ Adj R-squared = 0.3529
Total | 17878.875 199 89.843593 Root MSE = 7.6249
------------------------------------------------------------------------------
write | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
read | .5517051 .0527178 10.47 0.000 .4477445 .6556656
_cons | 23.95944 2.805744 8.54 0.000 18.42647 29.49242
------------------------------------------------------------------------------
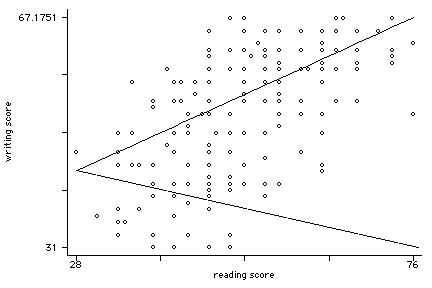
twoway (scatter write read, msym(oh))(lfit write read), legend(off)
regress write read female
Source | SS df MS Number of obs = 200
-------------+------------------------------ F( 2, 197) = 77.21
Model | 7856.32118 2 3928.16059 Prob > F = 0.0000
Residual | 10022.5538 197 50.8759077 R-squared = 0.4394
-------------+------------------------------ Adj R-squared = 0.4337
Total | 17878.875 199 89.843593 Root MSE = 7.1327
------------------------------------------------------------------------------
write | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
read | .5658869 .0493849 11.46 0.000 .468496 .6632778
female | 5.486894 1.014261 5.41 0.000 3.48669 7.487098
_cons | 20.22837 2.713756 7.45 0.000 14.87663 25.58011
------------------------------------------------------------------------------
predict p2
sort female p2
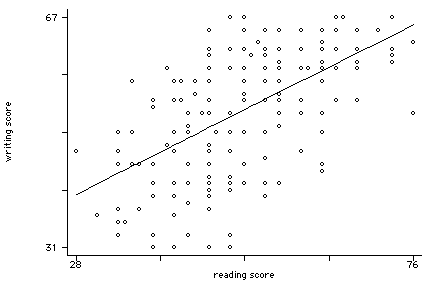
scatter write p2 read, msym(oh i) con(. L) sort
generate fxr = female*read
regress write c.read##i.female
Source | SS df MS Number of obs = 200
-------------+------------------------------ F( 3, 196) = 52.31
Model | 7949.6163 3 2649.8721 Prob > F = 0.0000
Residual | 9929.2587 196 50.6594831 R-squared = 0.4446
-------------+------------------------------ Adj R-squared = 0.4361
Total | 17878.875 199 89.843593 Root MSE = 7.1175
------------------------------------------------------------------------------
write | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
read | .6360156 .0714073 8.91 0.000 .4951904 .7768408
1.female | 12.49063 5.259266 2.37 0.019 2.118614 22.86265
|
female#|
c.read |
1 | -.133902 .0986707 -1.36 0.176 -.3284945 .0606905
|
_cons | 16.52388 3.845114 4.30 0.000 8.940769 24.10699
------------------------------------------------------------------------------
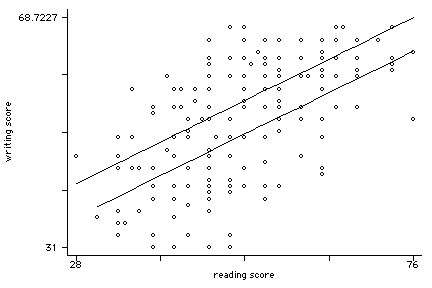
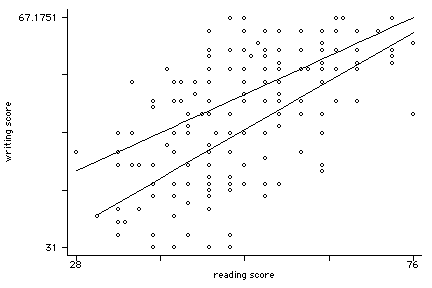
twoway (scatter write read, msym(oh)) (lfit write read if female==0) ///
(lfit write read if female==1), legend(off)xxxxxxx
Classical ANCOVA
- Partial out effects of one or more nuisance variables from the dependent variable.
- Random assignment to groups.
- One or more continuous variables.
- Statistical control vs experimental control.
Assumptions
- Independence
- Normality
- Homogeneity of variance
- Population within group regression coefficients are equal
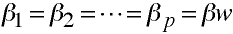
- covariate is measured without error
- Regression residuals are NID with mean 0 and equal variances
- Relationship between covariate & dependent variable is linear
- Covariate is related to dependent variable but is logically independent of treatment.
Selecting a Covariate
- The experiment contains one or more extraneous sources of variation--
- Related to the dependent variable
- Irrelevant to the independent variable
- Experimental control is not feasible or possible
- If possible measure covariate so that it does not include treatment effects
- Covariate obtained prior to treatment
- Covariate obtained after treatment but before it take effect
- Covariate is assumed to be unaffected by treatment
Logic of ANCOVA
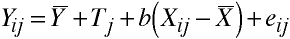
Which may be rewritten:
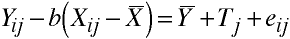
Homogeneity of Regression
- The previous two formulas involved the use of a common regression coefficient, b.
- Assumes that the slope of the regression line is constant across groups.
- Test the interaction of the independent variable and the covariate.
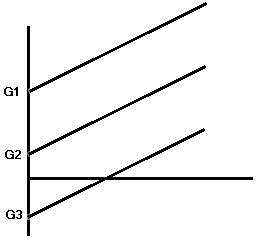
Steps in ANCOVA
- Include variable of scores on the dependent variable -- y
- Include variables of scores on the covariates -- c
- Create variables for coded group membership -- v
- Multiply each v by c to create interaction vectors -- cv
- Test variance added by step 4 -- homogeneity of regression slopes -- If n.s. procede to step 6; if sig. analyze as ATI
- Test variance added by step 3 -- If sig. then there are treatment effects, procede to step 7; if n.s. calculate regression with covariate only.
- If there are more than 2 levels perform multiple comparisons.
Numerical Example: coded using Effect Coding
input id y c1 c2 grp v1 v2 v3
1 6 1 6 1 1 0 0
2 9 1 7 1 1 0 0
3 8 2 15 1 1 0 0
4 8 3 13 1 1 0 0
5 12 3 18 1 1 0 0
6 12 4 9 1 1 0 0
7 10 4 16 1 1 0 0
8 8 5 10 1 1 0 0
9 12 5 16 1 1 0 0
10 13 6 18 1 1 0 0
11 13 4 12 2 0 1 0
12 16 4 12 2 0 1 0
13 15 5 17 2 0 1 0
14 16 6 9 2 0 1 0
15 19 6 20 2 0 1 0
16 17 8 18 2 0 1 0
17 19 8 16 2 0 1 0
18 23 9 20 2 0 1 0
19 19 10 10 2 0 1 0
20 22 10 17 2 0 1 0
21 20 7 8 3 0 0 1
22 22 7 14 3 0 0 1
23 24 9 11 3 0 0 1
24 26 9 11 3 0 0 1
25 24 10 16 3 0 0 1
26 25 11 20 3 0 0 1
27 28 11 19 3 0 0 1
28 27 12 19 3 0 0 1
29 29 13 12 3 0 0 1
30 26 13 16 3 0 0 1
31 27 7 16 4 -1 -1 -1
32 28 8 10 4 -1 -1 -1
33 25 8 13 4 -1 -1 -1
34 27 9 7 4 -1 -1 -1
35 31 9 15 4 -1 -1 -1
36 29 10 20 4 -1 -1 -1
37 32 10 16 4 -1 -1 -1
38 30 12 21 4 -1 -1 -1
39 32 12 15 4 -1 -1 -1
40 33 14 21 4 -1 -1 -1
end
/* using regress with factor variables */
regress y i.grp##c.c1
Source | SS df MS Number of obs = 40
-------------+------------------------------ F( 7, 32) = 109.27
Model | 2382.23359 7 340.319084 Prob > F = 0.0000
Residual | 99.6664092 32 3.11457529 R-squared = 0.9598
-------------+------------------------------ Adj R-squared = 0.9511
Total | 2481.9 39 63.6384615 Root MSE = 1.7648
------------------------------------------------------------------------------
y | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
grp |
2 | 3.435985 2.272924 1.51 0.140 -1.19381 8.06578
3 | 7.650473 3.069013 2.49 0.018 1.399098 13.90185
4 | 13.34207 3.017006 4.42 0.000 7.196633 19.48752
|
c1 | .9015152 .3434768 2.62 0.013 .2018757 1.601155
|
grp#c.c1 |
2 | .2026515 .4276252 0.47 0.639 -.6683925 1.073696
3 | .1489436 .4352144 0.34 0.734 -.7375591 1.035446
4 | .0402098 .4365514 0.09 0.927 -.8490164 .929436
|
_cons | 6.734848 1.29432 5.20 0.000 4.098405 9.371292
------------------------------------------------------------------------------
testparm grp#c.c1
( 1) 2.grp#c.c1 = 0
( 2) 3.grp#c.c1 = 0
( 3) 4.grp#c.c1 = 0
F( 3, 32) = 0.11
Prob > F = 0.9550
/* using anova */
anova y i.grp##c.c1
Number of obs = 40 R-squared = 0.9598
Root MSE = 1.76482 Adj R-squared = 0.9511
Source | Partial SS df MS F Prob > F
-----------+----------------------------------------------------
Model | 2382.23359 7 340.319084 109.27 0.0000
|
grp | 70.1635717 3 23.3878572 7.51 0.0006
c1 | 152.279387 1 152.279387 48.89 0.0000
grp#c1 | 1.00618243 3 .335394144 0.11 0.9550
|
Residual | 99.6664092 32 3.11457529
-----------+----------------------------------------------------
Total | 2481.9 39 63.6384615
Number of obs = 40 R-squared = 0.9598
Root MSE = 1.76482 Adj R-squared = 0.9511
Source | Partial SS df MS F Prob > F
-----------+----------------------------------------------------
Model | 2382.23359 7 340.319084 109.27 0.0000
|
grp | 70.1635717 3 23.3878572 7.51 0.0006
c1 | 152.279387 1 152.279387 48.89 0.0000
grp#c1 | 1.00618243 3 .335394144 0.11 0.9550
|
Residual | 99.6664092 32 3.11457529
-----------+----------------------------------------------------
Total | 2481.9 39 63.6384615
/* without interaction */
anova y i.grp c1
Number of obs = 40 R-squared = 0.9681
Root MSE = 1.85482 Adj R-squared = 0.9459
Source | Partial SS df MS F Prob > F
-----------+----------------------------------------------------
Model | 2402.77139 16 150.173212 43.65 0.0000
|
grp | 270.804721 3 90.2682404 26.24 0.0000
c1 | 186.671388 13 14.3593375 4.17 0.0014
|
Residual | 79.1286121 23 3.44037444
-----------+----------------------------------------------------
Total | 2481.9 39 63.6384615
/* back to regress with factor variables */
regress y i.grp c1
Source | SS df MS Number of obs = 40
-------------+------------------------------ F( 4, 35) = 206.97
Model | 2381.22741 4 595.306852 Prob > F = 0.0000
Residual | 100.672592 35 2.87635976 R-squared = 0.9594
-------------+------------------------------ Adj R-squared = 0.9548
Total | 2481.9 39 63.6384615 Root MSE = 1.696
------------------------------------------------------------------------------
y | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
grp |
2 | 4.453014 .8983061 4.96 0.000 2.629356 6.276673
3 | 8.411249 1.184014 7.10 0.000 6.007572 10.81493
4 | 13.01516 1.1535 11.28 0.000 10.67344 15.35689
|
c1 | 1.013052 .1337037 7.58 0.000 .7416185 1.284485
_cons | 6.355625 .703058 9.04 0.000 4.928341 7.782908
------------------------------------------------------------------------------
testparm i.grp
( 1) 2.grp = 0
( 2) 3.grp = 0
( 3) 4.grp = 0
F( 3, 35) = 48.19
Prob > F = 0.0000
/* compute original means */
table grp, contents(mean y)
----------+-----------
grp | mean(y)
----------+-----------
1 | 9.8
2 | 17.9
3 | 25.1
4 | 29.4
----------+-----------
/* compute adjusted means using margins */
/* margins will work with either regress or anova */
margins grp, asbalanced
Predictive margins Number of obs = 40
Model VCE : OLS
Expression : Linear prediction, predict()
------------------------------------------------------------------------------
| Delta-method
| Margin Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
grp |
1 | 14.08014 .7789391 18.08 0.000 12.55345 15.60684
2 | 18.53316 .5427882 34.14 0.000 17.46931 19.597
3 | 22.49139 .6373144 35.29 0.000 21.24228 23.74051
4 | 27.09531 .6165704 43.95 0.000 25.88685 28.30376
------------------------------------------------------------------------------
Regression Equation
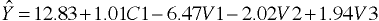
Separate Intercepts
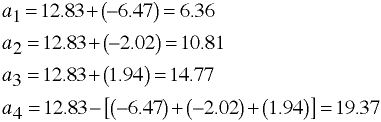
Computing Adjusted Means
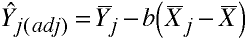
Multiple Covariates
- Effects of two or more covariates are removed from the dependent variable.
- Proceeds in a manner similar to the single covariate model.
Numerical Example
Same data as above example, except for the additional interaction terms:
/* run regression */
regress y c.c1##grp c.c2##grp
Source | SS df MS Number of obs = 40
-------------+------------------------------ F( 11, 28) = 71.35
Model | 2396.40189 11 217.854717 Prob > F = 0.0000
Residual | 85.4981125 28 3.05350402 R-squared = 0.9656
-------------+------------------------------ Adj R-squared = 0.9520
Total | 2481.9 39 63.6384615 Root MSE = 1.7474
------------------------------------------------------------------------------
y | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
c1 | .6228412 .4106184 1.52 0.141 -.2182726 1.463955
|
grp |
2 | 1.748814 3.135003 0.56 0.581 -4.672948 8.170576
3 | 9.113772 3.358697 2.71 0.011 2.233793 15.99375
4 | 14.68458 3.254535 4.51 0.000 8.017967 21.35119
|
grp#c.c1 |
2 | .3730242 .4858476 0.77 0.449 -.6221895 1.368238
3 | .4238656 .517841 0.82 0.420 -.6368836 1.484615
4 | .2512126 .5316271 0.47 0.640 -.8377761 1.340201
|
c2 | .1896132 .1565606 1.21 0.236 -.1310866 .5103131
|
grp#c.c2 |
2 | .0703098 .2157489 0.33 0.747 -.3716317 .5122513
3 | -.1858784 .2318612 -0.80 0.429 -.6608245 .2890676
4 | -.1372108 .2240572 -0.61 0.545 -.5961711 .3217495
|
_cons | 5.255291 1.770547 2.97 0.006 1.62849 8.882092
------------------------------------------------------------------------------
/* test homogeneity of regression slopes */
testparm grp#c.c1 grp#c.c2
( 1) 2.grp#c.c1 = 0
( 2) 3.grp#c.c1 = 0
( 3) 4.grp#c.c1 = 0
( 4) 2.grp#c.c2 = 0
( 5) 3.grp#c.c2 = 0
( 6) 4.grp#c.c2 = 0
F( 6, 28) = 0.43
Prob > F = 0.8553
/* using anova */
anova y c.c1##grp c.c2##grp
Number of obs = 40 R-squared = 0.9656
Root MSE = 1.74743 Adj R-squared = 0.9520
Source | Partial SS df MS F Prob > F
-----------+----------------------------------------------------
Model | 2396.40189 11 217.854717 71.35 0.0000
|
c1 | 85.0803048 1 85.0803048 27.86 0.0000
grp | 73.5988644 3 24.5329548 8.03 0.0005
grp#c1 | 2.40458727 3 .80152909 0.26 0.8518
c2 | 7.69362841 1 7.69362841 2.52 0.1237
grp#c2 | 5.14302215 3 1.71434072 0.56 0.6449
|
Residual | 85.4981125 28 3.05350402
-----------+----------------------------------------------------
Total | 2481.9 39 63.6384615
test grp#c.c1 grp#c.c2
Source | Partial SS df MS F Prob > F
--------------+----------------------------------------------------
grp#c1 grp#c2 | 7.80432184 6 1.30072031 0.43 0.8553
Residual | 85.4981125 28 3.05350402
/* anova without interaction */
anova y c.c1 c.c2 grp
Number of obs = 40 R-squared = 0.9624
Root MSE = 1.65656 Adj R-squared = 0.9569
Source | Partial SS df MS F Prob > F
-----------+----------------------------------------------------
Model | 2388.59757 5 477.719513 174.08 0.0000
|
c1 | 98.974038 1 98.974038 36.07 0.0000
c2 | 7.37015734 1 7.37015734 2.69 0.1105
grp | 420.189396 3 140.063132 51.04 0.0000
|
Residual | 93.3024343 34 2.74418925
-----------+----------------------------------------------------
Total | 2481.9 39 63.6384615
/* rerun as regression */
regress
Source | SS df MS Number of obs = 40
-------------+------------------------------ F( 5, 34) = 174.08
Model | 2388.59757 5 477.719513 Prob > F = 0.0000
Residual | 93.3024343 34 2.74418925 R-squared = 0.9624
-------------+------------------------------ Adj R-squared = 0.9569
Total | 2481.9 39 63.6384615 Root MSE = 1.6566
------------------------------------------------------------------------------
y | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
c1 | .8952525 .1490706 6.01 0.000 .5923047 1.1982
c2 | .1199612 .0731997 1.64 0.110 -.0287985 .2687209
|
grp |
2 | 4.60118 .8820702 5.22 0.000 2.808598 6.393762
3 | 8.996353 1.210347 7.43 0.000 6.536631 11.45607
4 | 13.46896 1.160214 11.61 0.000 11.11112 15.8268
|
_cons | 5.220638 .9753057 5.35 0.000 3.238579 7.202698
------------------------------------------------------------------------------
/* compute original means again */
table grp, contents(mean y)
----------+-----------
grp | mean(y)
----------+-----------
1 | 9.8
2 | 17.9
3 | 25.1
4 | 29.4
----------+-----------
/* compute adjusted means using margins */
margins grp, asbalanced
Predictive margins Number of obs = 40
Expression : Linear prediction, predict()
------------------------------------------------------------------------------
| Delta-method
| Margin Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
grp |
1 | 13.78338 .7820854 17.62 0.000 12.25052 15.31624
2 | 18.38456 .537869 34.18 0.000 17.33035 19.43876
3 | 22.77973 .646886 35.21 0.000 21.51186 24.0476
4 | 27.25234 .6098128 44.69 0.000 26.05713 28.44755
------------------------------------------------------------------------------
Regression Equation
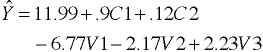
Separate Intercepts
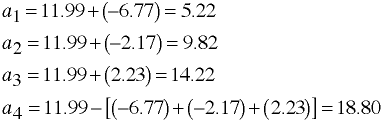
Interpretational Problems
- ANCOVA in experimental vs quasi-experimental research.
- "Equating" intact groups.
- "...what exactly does it mean to ask what the data would look like if they were not as they are" (Anderson, 1963).
- Lord, 1969-
- Controling for height of plant.
- What would be the yield of a 6ft variety be if they were 7ft?
- Depends on how you make them taller.
Specification Error
- Many problems in equating nonequivalent groups using ANCOVA are due to specification error.
- Can lead to over- or underadjustment of treatment means.
Extrapolation Errors
- When there is little or no overlap in group distributions
- Regression line for lowest group extrapolated upwards-
- Regression line for highest group extrapolated downwards.
Differential Growth
- Control using pretest as covariate
- Assumes rate of growth in nonequivalent groups are equal.
- Can lead to situations where differences in adjusted scores is due to differential growth.
Nonlinearity
- Do not assume that the regression is linear. Study the shape of the relationship. Avoiding erroneous assumptions and inappropriate analyses.
Measurement Error
- Measurement error in the covariate:
- Leads to underestimation of regression coefficient.
- This leads to underadjustment in the group lower on the CV.
- Often the treatment group in social intervention research.
Stata Example
These data are from a 1996 study (Gregoire, Kumar, Everitt, Henderson & Studd; also in Rabe-Hesketh & Everitt, 1999) on the efficacy of estrogen patches in treating postpartum depression. Women were randomly assigned to either a placebo control group (group=0, n=27) or estrogen patch group (group=1, n=34). Prior to the first treatment all patients took the Edinburgh Postnatal Depression Scale (EPDS). EPDS data was collected monthly for six months once the treatment began and average depression scores computed for each subject. Higher scores on the EDPS are indicative of higher levels of dsepression.
use http://www.philender.com/courses/data/depress1, clear
describe
Contains data from depress1.dta
obs: 61
vars: 4 18 Feb 2000 11:21
size: 1,220 (99.8% of memory free)
-------------------------------------------------------------------------------
1. subj float %9.0g
2. dep float %9.0g post-treatment depression score
3. pre float %9.0g pre-treatment depression score
4. group float %14.0g gl treatment group
-------------------------------------------------------------------------------
codebook group
group --------------------------------------------------------- treatment group
type: numeric (float)
label: gl
range: [0,1] units: 1
unique values: 2 coded missing: 0 / 61
tabulation: Freq. Numeric Label
27 0 placebo patch
34 1 estrogen patch
summarize pre dep
Variable | Obs Mean Std. Dev. Min Max
---------+-----------------------------------------------------
pre | 61 21.04033 3.722975 15 28
dep | 61 12.41284 5.407777 2 26.5
ttest pre, by(group)
Two-sample t test with equal variances
------------------------------------------------------------------------------
Group | Obs Mean Std. Err. Std. Dev. [95% Conf. Interval]
---------+--------------------------------------------------------------------
placebo | 27 20.77778 .7611158 3.954874 19.21328 22.34227
estrogen | 34 21.24882 .61301 3.574432 20.00165 22.496
---------+--------------------------------------------------------------------
combined | 61 21.04033 .476678 3.722975 20.08683 21.99383
---------+--------------------------------------------------------------------
diff | -.4710457 .9658499 -2.403707 1.461615
------------------------------------------------------------------------------
Degrees of freedom: 59
Ho: mean(placebo ) - mean(estrogen) = diff = 0
Ha: diff < 0 Ha: diff ~= 0 Ha: diff > 0
t = -0.4877 t = -0.4877 t = -0.4877
P < t = 0.3138 P > |t| = 0.6276 P > t = 0.6862
pwcorr dep pre, sig
| dep pre
-------------+------------------
dep | 1.0000
|
|
pre | 0.2920 1.0000
| 0.0224
|
/* a quick-and-dirty scatterplot */
plot dep pre
26.5 +
p | *
o |
s | *
t | *
- | *
t |
r | *
e | * * * * * *
a | * * *
t | * * * * * *
m | * *
e | * * *
n | * *
t | * * * * * *
| * *
d | * * * *
e | * * *
p | * * *
r | * * *
2 + * *
+----------------------------------------------------------------+
15 pre-treatment depression score 28
/* analysis without covariate */
regress dep i.group
Source | SS df MS Number of obs = 61
-------------+------------------------------ F( 1, 59) = 10.54
Model | 265.972224 1 265.972224 Prob > F = 0.0019
Residual | 1488.67078 59 25.2317081 R-squared = 0.1516
-------------+------------------------------ Adj R-squared = 0.1372
Total | 1754.643 60 29.2440501 Root MSE = 5.0231
------------------------------------------------------------------------------
dep | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
1.group | -4.20399 1.294842 -3.25 0.002 -6.794964 -1.613017
_cons | 14.75605 .9666994 15.26 0.000 12.82169 16.69041
------------------------------------------------------------------------------
margins group, asbalanced
Adjusted predictions Number of obs = 61
Model VCE : OLS
Expression : Linear prediction, predict()
------------------------------------------------------------------------------
| Delta-method
| Margin Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
group |
0 | 14.75605 .9666994 15.26 0.000 12.86135 16.65075
1 | 10.55206 .8614575 12.25 0.000 8.863633 12.24048
------------------------------------------------------------------------------
The ANCOVA
/* test for treat by covariate interation */
anova dep c.pre##group
Number of obs = 61 R-squared = 0.2541
Root MSE = 4.79187 Adj R-squared = 0.2148
Source | Partial SS df MS F Prob > F
-----------+----------------------------------------------------
Model | 445.80933 3 148.60311 6.47 0.0008
|
pre | 177.442758 1 177.442758 7.73 0.0074
group | 1.49167086 1 1.49167086 0.06 0.7997
group#pre | 3.19588284 1 3.19588284 0.14 0.7105
|
Residual | 1308.83367 57 22.9619943
-----------+----------------------------------------------------
Total | 1754.643 60 29.2440501
regress dep pre i.group
Source | SS df MS Number of obs = 61
-------------+------------------------------ F( 2, 58) = 9.78
Model | 442.613448 2 221.306724 Prob > F = 0.0002
Residual | 1312.02956 58 22.6211993 R-squared = 0.2523
-------------+------------------------------ Adj R-squared = 0.2265
Total | 1754.643 60 29.2440501 Root MSE = 4.7562
------------------------------------------------------------------------------
dep | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | .4618001 .1652592 2.79 0.007 .1309977 .7926024
1.group | -4.421519 1.2285 -3.60 0.001 -6.880629 -1.96241
_cons | 5.16087 3.553626 1.45 0.152 -1.952484 12.27422
------------------------------------------------------------------------------
table group, contents(mean dep)
---------------+-----------
treatment |
group | mean(dep)
---------------+-----------
placebo patch | 14.75605
estrogen patch | 10.55206
---------------+-----------
margins group, asbalanced
Predictive margins Number of obs = 61
Model VCE : OLS
Expression : Linear prediction, predict()
------------------------------------------------------------------------------
| Delta-method
| Margin Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
group |
0 | 14.87729 .9163541 16.24 0.000 13.08127 16.67332
1 | 10.45578 .8164047 12.81 0.000 8.855652 12.0559
------------------------------------------------------------------------------Interpretation
The interaction between the covariate (pre) and the treatment (group) was not significant, implying that we have homogeneity of regression slopes. In the final regression model, both the covariate and the treatment were statistically significant. Women with higher pretest scores on depression remain higher after treatment. Each point increase on the pretest was associated with about a .46 point increase on the predicted posttest score.
The effect of the estrogen patch was also significant. Women using the treatment patch had predicted depression scoress almost 4.5 points lower than women using the control patch.
Another Stata Example
These data examine a reading instruction program called "reading recovery." Students are randomly assigned to two treatment groups: a control group which receives standard reading instruction (treat = 0, n = 43) and the reading recovery group (treat = 1, n = 32).
There were two pretests administered at the beginning of the year. One test (pre1) consisted of dictation tasks, and the second (pre2) were early literacy skills. After four months of remedial reading instruction, the students we administered a standardized test of reading skills.
We will begin be examining the variables and determining if the treatment groups differ on the pretest measures.
use http://www.philender.com/courses/data/readexp, clear
describe
Contains data from readexp.dta
obs: 75
vars: 6 21 Dec 2000 21:29
size: 2,100 (99.8% of memory free)
-------------------------------------------------------------------------------
1. id float %9.0g
2. school float %9.0g
3. treat float %9.0g
4. pre1 float %9.0g
5. pre2 float %9.0g
6. post float %9.0g
-------------------------------------------------------------------------------
tabulate treat
treat | Freq. Percent Cum.
------------+-----------------------------------
0 | 43 57.33 57.33
1 | 32 42.67 100.00
------------+-----------------------------------
Total | 75 100.00
summarize pre1 pre2 post
Variable | Obs Mean Std. Dev. Min Max
---------+-----------------------------------------------------
pre1 | 75 8.44 7.571711 0 31
pre2 | 75 39.05333 18.32506 7 88
post | 75 33.26667 11.11528 12 64
corr pre1 pre2 post
(obs=75)
| pre1 pre2 post
---------+---------------------------
pre1 | 1.0000
pre2 | 0.6017 1.0000
post | 0.3202 0.5522 1.0000
stem pre1 if treat==0, lines(2)
Stem-and-leaf plot for pre1
0* | 00001222333344444
0. | 555667789
1* | 0134
1. | 568899
2* | 013
2. | 569
3* | 1
stem pre1 if treat==1, lines(2)
Stem-and-leaf plot for pre1
0* | 0001222223344
0. | 5556678889
1* | 000111
1. | 66
2* | 1
stem pre2 if treat==0, lines(1)
Stem-and-leaf plot for pre2
0* | 7
1* | 0668889
2* | 0013356899
3* | 0146789
4* | 13346
5* | 01225679
6* | 1366
7* |
8* | 8
stem pre2 if treat==1, lines(1)
Stem-and-leaf plot for pre2
0* | 9
1* | 367
2* | 12379
3* | 02358
4* | 001677
5* | 00123469
6* | 17
7* |
8* | 24
ttest pre1, by(treat)
Two-sample t test with equal variances
------------------------------------------------------------------------------
Group | Obs Mean Std. Err. Std. Dev. [95% Conf. Interval]
---------+--------------------------------------------------------------------
0 | 43 9.883721 1.337014 8.767389 7.185517 12.58192
1 | 32 6.5 .9002688 5.092689 4.66389 8.33611
---------+--------------------------------------------------------------------
combined | 75 8.44 .8743059 7.571711 6.697908 10.18209
---------+--------------------------------------------------------------------
diff | 3.383721 1.735173 -.0744738 6.841916
------------------------------------------------------------------------------
Degrees of freedom: 73
Ho: mean(0) - mean(1) = diff = 0
Ha: diff < 0 Ha: diff ~= 0 Ha: diff > 0
t = 1.9501 t = 1.9501 t = 1.9501
P < t = 0.9725 P > |t| = 0.0550 P > t = 0.0275
ttest pre2, by(treat)
Two-sample t test with equal variances
------------------------------------------------------------------------------
Group | Obs Mean Std. Err. Std. Dev. [95% Conf. Interval]
---------+--------------------------------------------------------------------
0 | 43 37.30233 2.769381 18.16005 31.71349 42.89116
1 | 32 41.40625 3.282671 18.56959 34.7112 48.1013
---------+--------------------------------------------------------------------
combined | 75 39.05333 2.115996 18.32506 34.83712 43.26955
---------+--------------------------------------------------------------------
diff | -4.103924 4.280596 -12.63514 4.427291
------------------------------------------------------------------------------
Degrees of freedom: 73
Ho: mean(0) - mean(1) = diff = 0
Ha: diff < 0 Ha: diff ~= 0 Ha: diff > 0
t = -0.9587 t = -0.9587 t = -0.9587
P < t = 0.1704 P > |t| = 0.3409 P > t = 0.8296
ranksum pre1, by(treat)
Two-sample Wilcoxon rank-sum (Mann-Whitney) test
treat | obs rank sum expected
---------+---------------------------------
0 | 43 1748 1634
1 | 32 1102 1216
---------+---------------------------------
combined | 75 2850 2850
unadjusted variance 8714.67
adjustment for ties -39.54
----------
adjusted variance 8675.12
Ho: pre1(treat==0) = pre1(treat==1)
z = 1.224
Prob > |z| = 0.2210
ranksum pre2, by(treat)
Two-sample Wilcoxon rank-sum (Mann-Whitney) test
treat | obs rank sum expected
---------+---------------------------------
0 | 43 1547.5 1634
1 | 32 1302.5 1216
---------+---------------------------------
combined | 75 2850 2850
unadjusted variance 8714.67
adjustment for ties -4.71
----------
adjusted variance 8709.96
Ho: pre2(treat==0) = pre2(treat==1)
z = -0.927
Prob > |z| = 0.3540
Interpretation
Because there appears to be a great deal of skewness in pre2 and some differences in the shapes of the distributions for pre1 Mann-Whitney tests (ranksum command) are preferable to the Student's t-tests for looking at pretest differences in the groups
Now, let's conduct the analysis of covariance.
/* without covariates */
regress post treat
Source | SS df MS Number of obs = 75
---------+------------------------------ F( 1, 73) = 8.39
Model | 942.050388 1 942.050388 Prob > F = 0.0050
Residual | 8200.61628 73 112.337209 R-squared = 0.1030
---------+------------------------------ Adj R-squared = 0.0908
Total | 9142.66667 74 123.54955 Root MSE = 10.599
------------------------------------------------------------------------------
post | Coef. Std. Err. t P>|t| [95% Conf. Interval]
---------+--------------------------------------------------------------------
treat | 7.165698 2.474476 2.896 0.005 2.234075 12.09732
_cons | 30.2093 1.616321 18.690 0.000 26.98798 33.43063
------------------------------------------------------------------------------
/* test treatment by slope interaction */
anova post c.pre1##treat c.pre2##treat
Number of obs = 75 R-squared = 0.3989
Root MSE = 8.92437 Adj R-squared = 0.3554
Source | Partial SS df MS F Prob > F
-----------+----------------------------------------------------
Model | 3647.21028 5 729.442056 9.16 0.0000
|
pre1 | .66873253 1 .66873253 0.01 0.9273
treat | 22.9971541 1 22.9971541 0.29 0.5928
treat#pre1 | 137.179391 1 137.179391 1.72 0.1937
pre2 | 1190.78252 1 1190.78252 14.95 0.0002
treat#pre2 | 144.906484 1 144.906484 1.82 0.1818
|
Residual | 5495.45639 69 79.6442955
-----------+----------------------------------------------------
Total | 9142.66667 74 123.54955
test treat#c.pre1 treat#c.pre2
Source | Partial SS df MS F Prob > F
----------------------+----------------------------------------------------
treat#pre1 treat#pre2 | 168.245679 2 84.1228394 1.06 0.3533
Residual | 5495.45639 69 79.6442955
/* the ancova */
regress post pre1 pre2 i.treat
Source | SS df MS Number of obs = 75
-------------+------------------------------ F( 3, 71) = 14.54
Model | 3478.9646 3 1159.65487 Prob > F = 0.0000
Residual | 5663.70207 71 79.7704516 R-squared = 0.3805
-------------+------------------------------ Adj R-squared = 0.3543
Total | 9142.66667 74 123.54955 Root MSE = 8.9314
------------------------------------------------------------------------------
post | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre1 | .1698471 .1843945 0.92 0.360 -.1978251 .5375193
pre2 | .2726797 .0747458 3.65 0.001 .1236409 .4217185
1.treat | 6.621356 2.253639 2.94 0.004 2.127727 11.11499
_cons | 18.35899 2.525568 7.27 0.000 13.32315 23.39483
------------------------------------------------------------------------------
regress post pre2 i.treat
Source | SS df MS Number of obs = 75
-------------+------------------------------ F( 2, 72) = 21.43
Model | 3411.28428 2 1705.64214 Prob > F = 0.0000
Residual | 5731.38239 72 79.6025332 R-squared = 0.3731
-------------+------------------------------ Adj R-squared = 0.3557
Total | 9142.66667 74 123.54955 Root MSE = 8.922
------------------------------------------------------------------------------
post | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre2 | .3172027 .0569533 5.57 0.000 .2036683 .4307371
1.treat | 5.863922 2.096051 2.80 0.007 1.68552 10.04232
_cons | 18.3769 2.522833 7.28 0.000 13.34773 23.40608
------------------------------------------------------------------------------
/* unadjusted means */
tabstat post, by(treat)
Summary for variables: post
by categories of: treat
treat | mean
---------+----------
0 | 30.2093
1 | 37.375
---------+----------
Total | 33.26667
--------------------
margins treat, asbalanced
Predictive margins Number of obs = 75
Model VCE : OLS
Expression : Linear prediction, predict()
------------------------------------------------------------------------------
| Delta-method
| Margin Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
treat |
0 | 30.76473 1.364246 22.55 0.000 28.09085 33.4386
1 | 36.62865 1.582889 23.14 0.000 33.52624 39.73105
------------------------------------------------------------------------------Interpretation
The interaction between the covariates (pre1 & pre2) and the treatment (treat) were not significant, implying that we have homogeneity of regression slopes. In the regression model with no interactions pre1 was not significant and was dropped from the analysis. Both the covariate (pre2) and the treatment were statistically significant. Students with higher pretest scores on tended to have higher posttest scores. Each point increase on the pre2 was associated with about a .32 point increase on the predicted posttest score.
The effect of the reading recovery was also significant while controling for the initial level of pre2. Students receiving the treatment had predicted posttest scores almost 5.9 points higher than students in the control group. The predicted change without the covariate was approximately 7.2 points. Including the covariate reduced the amount of predicted change.
Linear Statistical Models Course
Phil Ender, 24sep10, 22Feb00