

In this unit we will cover several logistic regression models along with some diagnostic techniques.
Example 1
use http://www.philender.com/courses/data/honors, clear
xi: logit honors lang math female i.ses, nolog
i.ses _Ises_1-3 (naturally coded; _Ises_1 omitted)
Logit estimates Number of obs = 200
LR chi2(5) = 87.30
Prob > chi2 = 0.0000
Log likelihood = -71.994756 Pseudo R2 = 0.3774
------------------------------------------------------------------------------
honors | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
lang | .0687277 .0287044 2.39 0.017 .0124681 .1249873
math | .1358904 .0336874 4.03 0.000 .0698642 .2019166
female | 1.145726 .4513589 2.54 0.011 .2610792 2.030374
_Ises_2 | -1.040402 .5791511 -1.80 0.072 -2.175517 .094713
_Ises_3 | .0541296 .5945439 0.09 0.927 -1.111155 1.219414
_cons | -12.55332 1.838493 -6.83 0.000 -16.1567 -8.949939
------------------------------------------------------------------------------
test _Ises_2 _Ises_3
( 1) _Ises_2 = 0
( 2) _Ises_3 = 0
chi2( 2) = 6.13
Prob > chi2 = 0.0466
linktest, nolog /* general test of model specification */
Logit estimates Number of obs = 200
LR chi2(2) = 87.39
Prob > chi2 = 0.0000
Log likelihood = -71.950384 Pseudo R2 = 0.3778
------------------------------------------------------------------------------
honors | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
_hat | .9703938 .172421 5.63 0.000 .6324549 1.308333
_hatsq | -.0236258 .0801491 -0.29 0.768 -.1807151 .1334635
_cons | .041176 .2706831 0.15 0.879 -.4893531 .5717051
------------------------------------------------------------------------------
predict pprob /* predicted probabilities */
predict r, resid /* pearson residuals */
predict h, hat /* leverage*/
predict db, dbeta /* pregibon dbeta */
predict dx2, dx2 /* hosmer & lemeshow influence */
scatter h r, xline(0) msym(Oh) jitter(2)
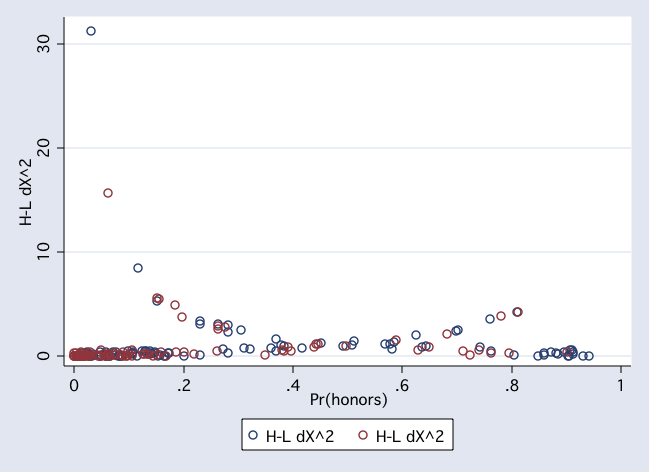
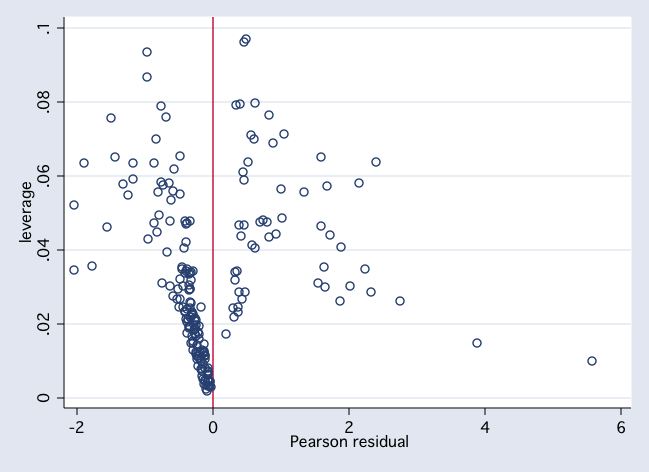 twoway (scatter dx2 pprob if female, msym(Oh) jitter(2)) ///
(scatter dx2 pprob if ~female, msym(Oh) jitter(2))
twoway (scatter dx2 pprob if female, msym(Oh) jitter(2)) ///
(scatter dx2 pprob if ~female, msym(Oh) jitter(2))
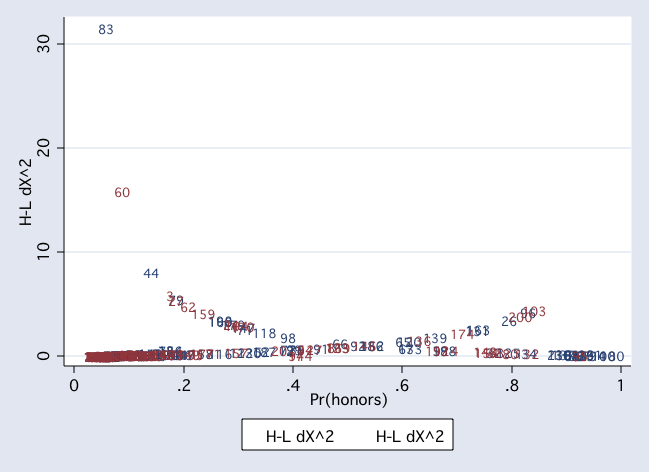
 twoway (scatter dx2 pprob if female, msym(i) mlab(id) jitter(2)) ///
(scatter dx2 pprob if ~female, msym(i) mlab(id) jitter(2))
twoway (scatter dx2 pprob if female, msym(i) mlab(id) jitter(2)) ///
(scatter dx2 pprob if ~female, msym(i) mlab(id) jitter(2))
 scatter dx2 pprob if female [w=db], msym(Oh) jitter(2)
scatter dx2 pprob if female [w=db], msym(Oh) jitter(2)
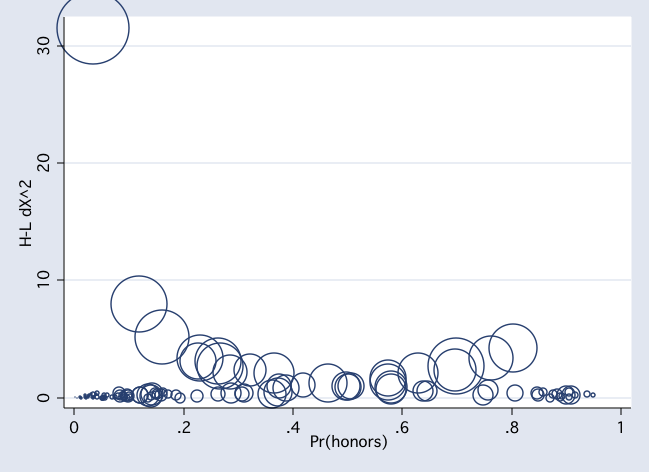 scatter dx2 pprob if ~female [w=db], msym(Oh) jitter(2)
scatter dx2 pprob if ~female [w=db], msym(Oh) jitter(2)
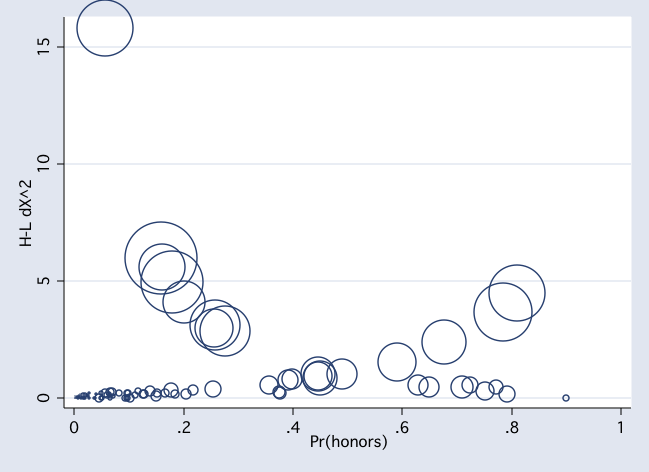
Example 2
use http://www.philender.com/courses/data/apilog, clear
logit hiqual yr_rnd meals cred_ml, nolog
Logit estimates Number of obs = 707
LR chi2(3) = 385.27
Prob > chi2 = 0.0000
Log likelihood = -156.38516 Pseudo R2 = 0.5519
------------------------------------------------------------------------------
hiqual | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
yr_rnd | -1.185658 .50163 -2.36 0.018 -2.168835 -.2024813
meals | -.0932877 .0084252 -11.07 0.000 -.1098008 -.0767746
cred_ml | .7415145 .3152036 2.35 0.019 .1237268 1.359302
_cons | 2.411226 .3987573 6.05 0.000 1.629676 3.192776
------------------------------------------------------------------------------
linktest, nolog
Logit estimates Number of obs = 707
LR chi2(2) = 391.76
Prob > chi2 = 0.0000
Log likelihood = -153.13783 Pseudo R2 = 0.5612
------------------------------------------------------------------------------
hiqual | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
_hat | 1.209837 .1280197 9.45 0.000 .9589229 1.460751
_hatsq | .0735317 .026548 2.77 0.006 .0214986 .1255648
_cons | -.1381412 .1636431 -0.84 0.399 -.4588757 .1825933
------------------------------------------------------------------------------
generate ym=yr_rnd*meals
logit hiqual yr_rnd meals cred_ml ym, nolog
Logit estimates Number of obs = 707
LR chi2(4) = 390.13
Prob > chi2 = 0.0000
Log likelihood = -153.95333 Pseudo R2 = 0.5589
------------------------------------------------------------------------------
hiqual | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
yr_rnd | -2.816989 .8625011 -3.27 0.001 -4.50746 -1.126518
meals | -.1014958 .0098204 -10.34 0.000 -.1207434 -.0822483
cred_ml | .7795476 .3205748 2.43 0.015 .1512326 1.407863
ym | .0459029 .0188068 2.44 0.015 .0090423 .0827635
_cons | 2.668048 .429688 6.21 0.000 1.825875 3.510221
------------------------------------------------------------------------------
linktest, nolog
Logit estimates Number of obs = 707
LR chi2(2) = 390.87
Prob > chi2 = 0.0000
Log likelihood = -153.58393 Pseudo R2 = 0.5600
------------------------------------------------------------------------------
hiqual | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
_hat | 1.063142 .1154731 9.21 0.000 .8368188 1.289465
_hatsq | .0279257 .031847 0.88 0.381 -.0344934 .0903447
_cons | -.0605556 .1684181 -0.36 0.719 -.3906491 .2695378
------------------------------------------------------------------------------
Example 3
use http://www.philender.com/courses/data/apilog, clear
summarize full, meanonly
generate fullc=full-r(mean)
generate yxfc=yr_rnd*fullc
logit hiqual avg_ed yr_rnd meals fullc yxfc, nolog
Logit estimates Number of obs = 1158
LR chi2(5) = 933.71
Prob > chi2 = 0.0000
Log likelihood = -263.83452 Pseudo R2 = 0.6389
------------------------------------------------------------------------------
hiqual | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
avg_ed | 1.968948 .2850136 6.91 0.000 1.410332 2.527564
yr_rnd | -.5484941 .3680305 -1.49 0.136 -1.269821 .1728325
meals | -.0789775 .0079544 -9.93 0.000 -.0945677 -.0633872
fullc | .0499983 .01452 3.44 0.001 .0215397 .0784569
yxfc | -.1329371 .0325101 -4.09 0.000 -.1966557 -.0692185
_cons | -3.655163 1.016972 -3.59 0.000 -5.648392 -1.661935
------------------------------------------------------------------------------
logit, or
Logit estimates Number of obs = 1158
LR chi2(5) = 933.71
Prob > chi2 = 0.0000
Log likelihood = -263.83452 Pseudo R2 = 0.6389
------------------------------------------------------------------------------
hiqual | Odds Ratio Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
avg_ed | 7.163138 2.041592 6.91 0.000 4.097315 12.52297
yr_rnd | .5778193 .2126551 -1.49 0.136 .280882 1.188667
meals | .9240607 .0073503 -9.93 0.000 .9097661 .93858
fullc | 1.051269 .0152644 3.44 0.001 1.021773 1.081617
yxfc | .8755202 .0284632 -4.09 0.000 .8214734 .9331228
------------------------------------------------------------------------------
predict p1
predict stdres, rstand /* standardized Pearson residual (adj. for # sharing covariate pattern) */
predict dv, dev /* deviance residual */
predict hat, hat /* leverage */
predict dx2, dx2 /* Hosmer and Lemeshow Delta chi-squared influence statistic */
predict dd, dd /* Hosmer and Lemeshow Delta-D influence statistic */
scatter stdres p1, mlab(snum) msym(i) yline(0)
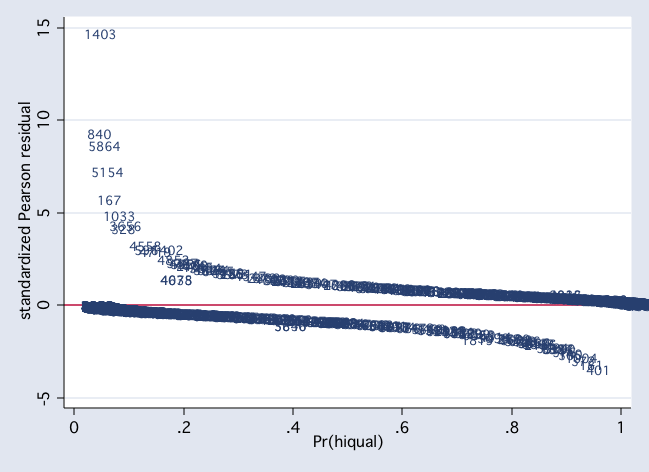 scatter stdres snum, mlab(snum) msym(i) yline(0)
scatter stdres snum, mlab(snum) msym(i) yline(0)
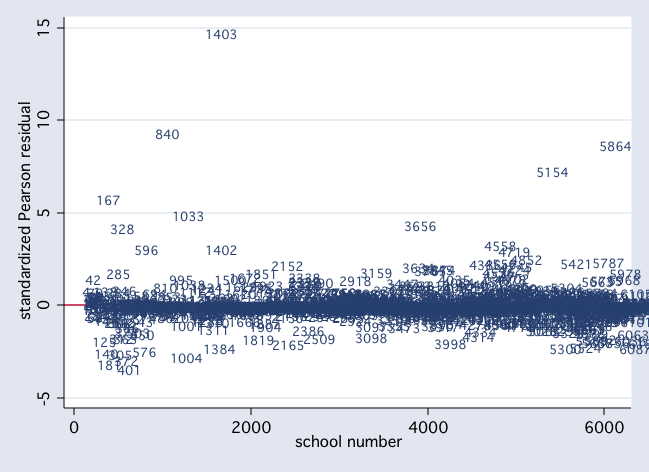 scatter dv p1, mlab(snum) msym(i) yline(0)
scatter dv p1, mlab(snum) msym(i) yline(0)
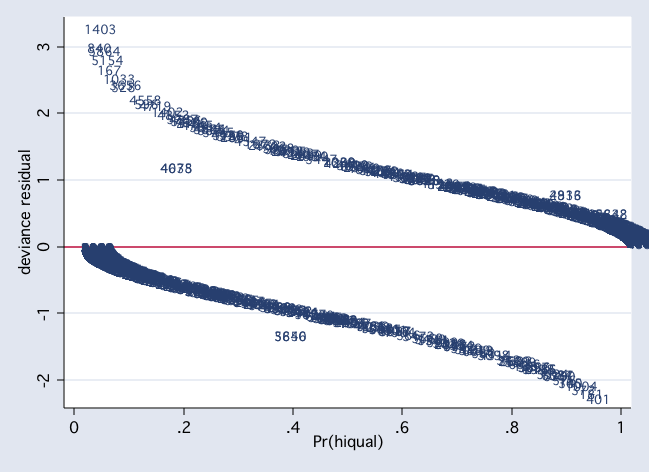 scatter hat p1, mlab(snum) msym(i) yline(0)
scatter hat p1, mlab(snum) msym(i) yline(0)
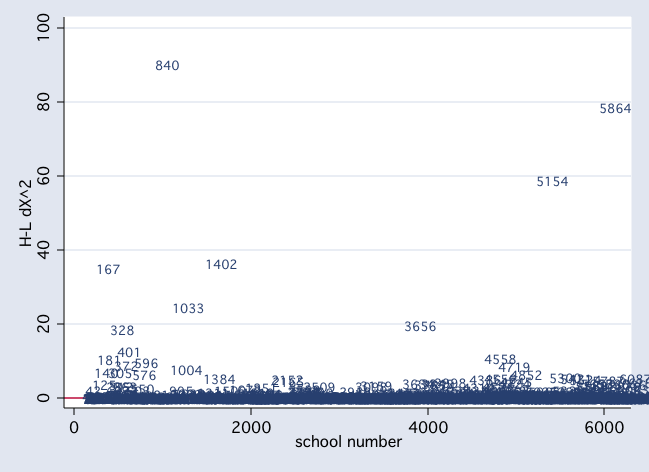
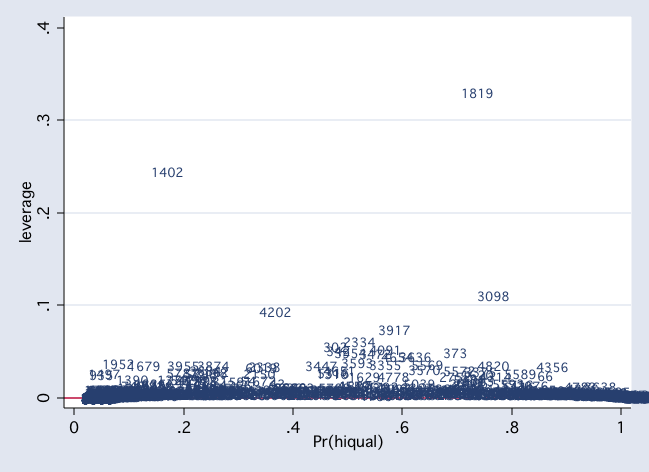 scatter dx2 snum, mlab(snum) msym(i) yline(0)
scatter dx2 snum, mlab(snum) msym(i) yline(0)
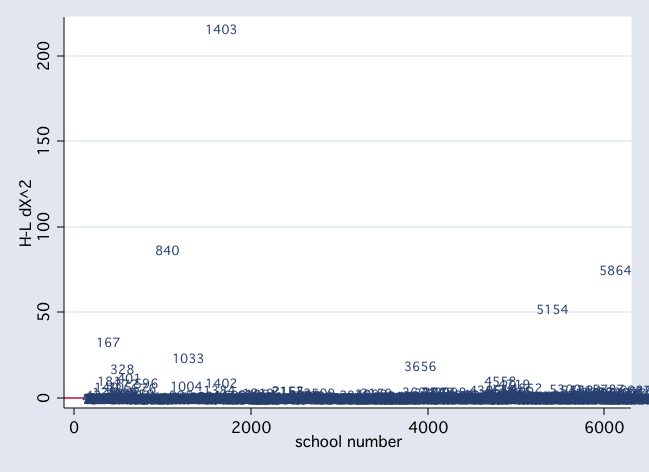 scatter dd snum, mlab(snum) msym(i) yline(0)
scatter dd snum, mlab(snum) msym(i) yline(0)
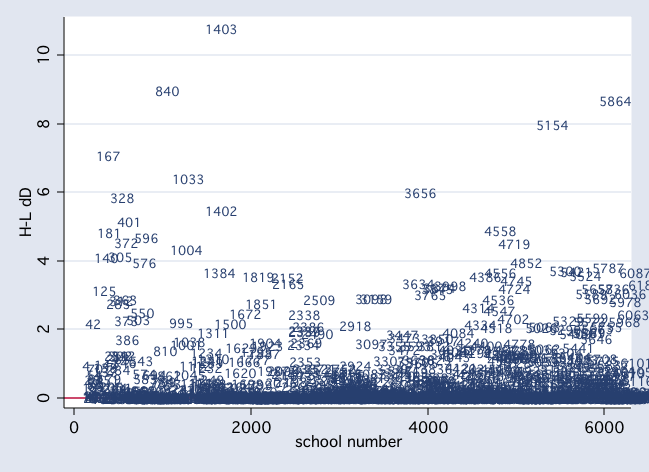 drop if snum==1403
logit hiqual avg_ed yr_rnd meals fullc yxfc, nolog
Logit estimates Number of obs = 1157
LR chi2(5) = 943.15
Prob > chi2 = 0.0000
Log likelihood = -257.99083 Pseudo R2 = 0.6464
------------------------------------------------------------------------------
hiqual | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
avg_ed | 2.030088 .2915102 6.96 0.000 1.458739 2.601437
yr_rnd | -.7044717 .3864407 -1.82 0.068 -1.461882 .0529381
meals | -.0797143 .0080847 -9.86 0.000 -.0955601 -.0638686
fullc | .0504368 .0146263 3.45 0.001 .0217697 .0791038
yxfc | -.1078501 .0372207 -2.90 0.004 -.1808013 -.034899
_cons | -3.819562 1.035962 -3.69 0.000 -5.850011 -1.789114
------------------------------------------------------------------------------
predict p2
predict stdres2, rstand
predict dx22, dx2
scatter stdres2 p2, mlab(snum) msym(i) yline(0)
drop if snum==1403
logit hiqual avg_ed yr_rnd meals fullc yxfc, nolog
Logit estimates Number of obs = 1157
LR chi2(5) = 943.15
Prob > chi2 = 0.0000
Log likelihood = -257.99083 Pseudo R2 = 0.6464
------------------------------------------------------------------------------
hiqual | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
avg_ed | 2.030088 .2915102 6.96 0.000 1.458739 2.601437
yr_rnd | -.7044717 .3864407 -1.82 0.068 -1.461882 .0529381
meals | -.0797143 .0080847 -9.86 0.000 -.0955601 -.0638686
fullc | .0504368 .0146263 3.45 0.001 .0217697 .0791038
yxfc | -.1078501 .0372207 -2.90 0.004 -.1808013 -.034899
_cons | -3.819562 1.035962 -3.69 0.000 -5.850011 -1.789114
------------------------------------------------------------------------------
predict p2
predict stdres2, rstand
predict dx22, dx2
scatter stdres2 p2, mlab(snum) msym(i) yline(0)
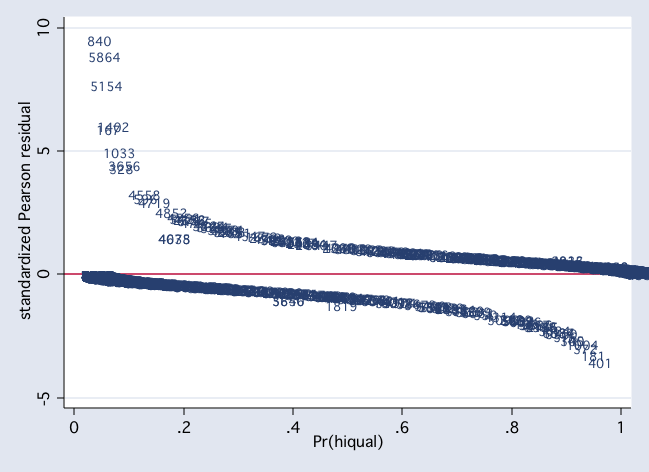 scatter dx22 snum, mlab(snum) msym(i) yline(0)
scatter dx22 snum, mlab(snum) msym(i) yline(0)
Categorical Data Analysis Course
Phil Ender