

Example with Binary Response Variable
The binary response example is derived from the previous example by converting depression scores to 0/1 values with a cut point of 11 and retaining only the 61 observations with complete data.
use http://www.gseis.ucla.edu/courses/data/deprl, clear
list in 1/3, nodisplay noobs nolabel /* output edited */
id dep1 dep2 dep3 dep4 dep5 dep6 treat pre
1 1 1 1 1 1 1 0 18
2 1 1 1 1 1 0 0 27
3 1 1 1 0 0 0 0 16
summarize
Variable | Obs Mean Std. Dev. Min Max
-------------+-----------------------------------------------------
id | 61 31 17.75293 1 61
dep1 | 61 .7213115 .4520748 0 1
dep2 | 61 .6885246 .4669398 0 1
dep3 | 61 .5409836 .502453 0 1
dep4 | 61 .4754098 .5035394 0 1
dep5 | 61 .3770492 .4886694 0 1
dep6 | 61 .2459016 .4341942 0 1
treat | 61 .557377 .500819 0 1
pre | 61 21.03279 3.710199 15 28
tab1 dep1 dep2 dep3 dep4 dep5 dep6 treat
-> tabulation of dep1
1 dep | Freq. Percent Cum.
------------+-----------------------------------
0 | 17 27.87 27.87
1 | 44 72.13 100.00
------------+-----------------------------------
Total | 61 100.00
-> tabulation of dep2
2 dep | Freq. Percent Cum.
------------+-----------------------------------
0 | 19 31.15 31.15
1 | 42 68.85 100.00
------------+-----------------------------------
Total | 61 100.00
-> tabulation of dep3
3 dep | Freq. Percent Cum.
------------+-----------------------------------
0 | 28 45.90 45.90
1 | 33 54.10 100.00
------------+-----------------------------------
Total | 61 100.00
-> tabulation of dep4
4 dep | Freq. Percent Cum.
------------+-----------------------------------
0 | 32 52.46 52.46
1 | 29 47.54 100.00
------------+-----------------------------------
Total | 61 100.00
-> tabulation of dep5
5 dep | Freq. Percent Cum.
------------+-----------------------------------
0 | 38 62.30 62.30
1 | 23 37.70 100.00
------------+-----------------------------------
Total | 61 100.00
-> tabulation of dep6
6 dep | Freq. Percent Cum.
------------+-----------------------------------
0 | 46 75.41 75.41
1 | 15 24.59 100.00
------------+-----------------------------------
Total | 61 100.00
-> tabulation of treat
treat | Freq. Percent Cum.
------------+-----------------------------------
placebo | 27 44.26 44.26
estrogen | 34 55.74 100.00
------------+-----------------------------------
Total | 61 100.00
corr dep1 dep2 dep3 dep4 dep5 dep6
(obs=61)
| dep1 dep2 dep3 dep4 dep5 dep6
-------------+------------------------------------------------------
dep1 | 1.0000
dep2 | 0.4504 1.0000
dep3 | 0.3079 0.6591 1.0000
dep4 | 0.2256 0.4985 0.6134 1.0000
dep5 | 0.2573 0.5233 0.5130 0.7495 1.0000
dep6 | 0.1851 0.3019 0.5260 0.5236 0.6554 1.0000
reshape long dep, i(id) j(visit)
(note: j = 1 2 3 4 5 6)
Data wide -> long
-----------------------------------------------------------------------------
Number of obs. 61 -> 366
Number of variables 9 -> 5
j variable (6 values) -> visit
xij variables:
dep1 dep2 ... dep6 -> dep
-----------------------------------------------------------------------------
list in 1/18, nolabel
id visit dep treat pre
1. 1 1 1 0 18
2. 1 2 1 0 18
3. 1 3 1 0 18
4. 1 4 1 0 18
5. 1 5 1 0 18
6. 1 6 1 0 18
7. 2 1 1 0 27
8. 2 2 1 0 27
9. 2 3 1 0 27
10. 2 4 1 0 27
11. 2 5 1 0 27
12. 2 6 0 0 27
13. 3 1 1 0 16
14. 3 2 1 0 16
15. 3 3 1 0 16
16. 3 4 0 0 16
17. 3 5 0 0 16
18. 3 6 0 0 16
logit dep pre treat
Logit estimates Number of obs = 366
LR chi2(2) = 67.12
Prob > chi2 = 0.0000
Log likelihood = -220.08461 Pseudo R2 = 0.1323
------------------------------------------------------------------------------
dep | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | .1672391 .0337122 4.96 0.000 .1011644 .2333139
treat | -1.573125 .2415083 -6.51 0.000 -2.046473 -1.099778
_cons | -2.586276 .6907273 -3.74 0.000 -3.940077 -1.232476
------------------------------------------------------------------------------
xtgee dep pre treat, i(id) link(logit) fam(bin) corr(ind)
GEE population-averaged model Number of obs = 366
Group variable: id Number of groups = 61
Link: logit Obs per group: min = 6
Family: binomial avg = 6.0
Correlation: independent max = 6
Wald chi2(2) = 53.47
Scale parameter: 1 Prob > chi2 = 0.0000
Pearson chi2(366): 369.76 Deviance = 440.17
Dispersion (Pearson): 1.010272 Dispersion = 1.202648
------------------------------------------------------------------------------
dep | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | .1672391 .0337125 4.96 0.000 .1011638 .2333145
treat | -1.573125 .2415102 -6.51 0.000 -2.046476 -1.099774
_cons | -2.586276 .6907322 -3.74 0.000 -3.940087 -1.232466
------------------------------------------------------------------------------
xtcorr
Estimated within-id correlation matrix R:
c1 c2 c3 c4 c5 c6
r1 1.0000
r2 0.0000 1.0000
r3 0.0000 0.0000 1.0000
r4 0.0000 0.0000 0.0000 1.0000
r5 0.0000 0.0000 0.0000 0.0000 1.0000
r6 0.0000 0.0000 0.0000 0.0000 0.0000 1.0000
xtgee dep pre treat, i(id) link(logit) fam(bin) corr(exc)
GEE population-averaged model Number of obs = 366
Group variable: id Number of groups = 61
Link: logit Obs per group: min = 6
Family: binomial avg = 6.0
Correlation: exchangeable max = 6
Wald chi2(2) = 23.63
Scale parameter: 1 Prob > chi2 = 0.0000
------------------------------------------------------------------------------
dep | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | .1672391 .0507097 3.30 0.001 .0678499 .2666284
treat | -1.573125 .3632751 -4.33 0.000 -2.285131 -.8611189
_cons | -2.586276 1.038986 -2.49 0.013 -4.622652 -.5499009
------------------------------------------------------------------------------
xtcorr
Estimated within-id correlation matrix R:
c1 c2 c3 c4 c5 c6
r1 1.0000
r2 0.2525 1.0000
r3 0.2525 0.2525 1.0000
r4 0.2525 0.2525 0.2525 1.0000
r5 0.2525 0.2525 0.2525 0.2525 1.0000
r6 0.2525 0.2525 0.2525 0.2525 0.2525 1.0000
generate pxt = pre*treat
xtgee dep pre treat pxt, i(id) link(logit) fam(bin) corr(ar1) t(visit)
GEE population-averaged model Number of obs = 366
Group and time vars: id visit Number of groups = 61
Link: logit Obs per group: min = 6
Family: binomial avg = 6.0
Correlation: AR(1) max = 6
Wald chi2(3) = 19.59
Scale parameter: 1 Prob > chi2 = 0.0002
------------------------------------------------------------------------------
dep | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | .1526517 .0748497 2.04 0.041 .0059491 .2993544
treat | -.9262238 2.107262 -0.44 0.660 -5.056382 3.203935
pxt | -.0245282 .1003177 -0.24 0.807 -.2211473 .1720909
_cons | -2.378658 1.516685 -1.57 0.117 -5.351307 .5939899
------------------------------------------------------------------------------
xtgee dep pre treat, i(id) link(logit) fam(bin) corr(ar1) t(visit)
GEE population-averaged model Number of obs = 366
Group and time vars: id visit Number of groups = 61
Link: logit Obs per group: min = 6
Family: binomial avg = 6.0
Correlation: AR(1) max = 6
Wald chi2(2) = 19.71
Scale parameter: 1 Prob > chi2 = 0.0000
------------------------------------------------------------------------------
dep | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | .1390761 .049729 2.80 0.005 .041609 .2365432
treat | -1.434432 .359136 -3.99 0.000 -2.138326 -.7305387
_cons | -2.107566 1.030142 -2.05 0.041 -4.126608 -.0885242
------------------------------------------------------------------------------
xtcorr
Estimated within-id correlation matrix R:
c1 c2 c3 c4 c5 c6
r1 1.0000
r2 0.5256 1.0000
r3 0.2762 0.5256 1.0000
r4 0.1452 0.2762 0.5256 1.0000
r5 0.0763 0.1452 0.2762 0.5256 1.0000
r6 0.0401 0.0763 0.1452 0.2762 0.5256 1.0000
xtgee, eform
GEE population-averaged model Number of obs = 366
Group and time vars: id visit Number of groups = 61
Link: logit Obs per group: min = 6
Family: binomial avg = 6.0
Correlation: AR(1) max = 6
Wald chi2(2) = 19.71
Scale parameter: 1 Prob > chi2 = 0.0000
------------------------------------------------------------------------------
dep | Odds Ratio Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | 1.149212 .0571492 2.80 0.005 1.042487 1.266862
treat | .2382506 .0855644 -3.99 0.000 .1178519 .4816495
------------------------------------------------------------------------------
xi: xtgee dep pre treat i.visit, i(id) link(logit) fam(bin) corr(ar1) t(visit)
GEE population-averaged model Number of obs = 366
Group and time vars: id visit Number of groups = 61
Link: logit Obs per group: min = 6
Family: binomial avg = 6.0
Correlation: AR(1) max = 6
Wald chi2(7) = 43.18
Scale parameter: 1 Prob > chi2 = 0.0000
------------------------------------------------------------------------------
dep | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | .1663743 .0534293 3.11 0.002 .0616548 .2710938
treat | -1.736828 .3977053 -4.37 0.000 -2.516316 -.9573399
_Ivisit_2 | -.1606584 .3089872 -0.52 0.603 -.7662623 .4449455
_Ivisit_3 | -.9535964 .3704544 -2.57 0.010 -1.679674 -.2275192
_Ivisit_4 | -1.301396 .4028895 -3.23 0.001 -2.091045 -.5117472
_Ivisit_5 | -1.806927 .4283058 -4.22 0.000 -2.646391 -.9674631
_Ivisit_6 | -2.567095 .4682141 -5.48 0.000 -3.484778 -1.649412
_cons | -1.335994 1.104602 -1.21 0.226 -3.500974 .8289849
------------------------------------------------------------------------------
xtgee dep pre treat visit, i(id) link(logit) fam(bin) corr(ar1) t(visit)
GEE population-averaged model Number of obs = 366
Group and time vars: id visit Number of groups = 61
Link: logit Obs per group: min = 6
Family: binomial avg = 6.0
Correlation: AR(1) max = 6
Wald chi2(3) = 42.84
Scale parameter: 1 Prob > chi2 = 0.0000
------------------------------------------------------------------------------
dep | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | .1756632 .0533018 3.30 0.001 .0711937 .2801328
treat | -1.759441 .3946169 -4.46 0.000 -2.532876 -.9860058
visit | -.5189469 .0917666 -5.66 0.000 -.6988061 -.3390876
_cons | -.8539732 1.095211 -0.78 0.436 -3.000546 1.2926
------------------------------------------------------------------------------
/* test visit categorical versus visit continuous */
xtgee dep pre treat visit _Ivisit_3 _Ivisit_4 _Ivisit_5 _Ivisit_6, i(id) link(logit) fam(bin) corr(ar1) t(visit)
GEE population-averaged model Number of obs = 366
Group and time vars: id visit Number of groups = 61
Link: logit Obs per group: min = 6
Family: binomial avg = 6.0
Correlation: AR(1) max = 6
Wald chi2(7) = 43.18
Scale parameter: 1 Prob > chi2 = 0.0000
------------------------------------------------------------------------------
dep | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | .1663743 .0534293 3.11 0.002 .0616548 .2710938
treat | -1.736828 .3977053 -4.37 0.000 -2.516316 -.9573399
visit | -.1606584 .3089872 -0.52 0.603 -.7662623 .4449455
_Ivisit_3 | -.6322796 .4825321 -1.31 0.190 -1.578025 .3134658
_Ivisit_4 | -.8194209 .8082586 -1.01 0.311 -2.403579 .764737
_Ivisit_5 | -1.164294 1.121156 -1.04 0.299 -3.361719 1.033132
_Ivisit_6 | -1.763803 1.434124 -1.23 0.219 -4.574634 1.047028
_cons | -1.175336 1.185178 -0.99 0.321 -3.498242 1.14757
------------------------------------------------------------------------------
test _Ivisit_3 _Ivisit_4 _Ivisit_5 _Ivisit_6
( 1) _Ivisit_3 = 0
( 2) _Ivisit_4 = 0
( 3) _Ivisit_5 = 0
( 4) _Ivisit_6 = 0
chi2( 4) = 2.58
Prob > chi2 = 0.6303
/* rerun model with continuous time */
xtgee dep pre treat visit, i(id) link(logit) fam(bin) corr(ar1) t(visit)
Iteration 1: tolerance = .14672104
Iteration 2: tolerance = .00108396
Iteration 3: tolerance = .00009351
Iteration 4: tolerance = 3.269e-06
Iteration 5: tolerance = 2.043e-07
GEE population-averaged model Number of obs = 366
Group and time vars: id visit Number of groups = 61
Link: logit Obs per group: min = 6
Family: binomial avg = 6.0
Correlation: AR(1) max = 6
Wald chi2(3) = 42.84
Scale parameter: 1 Prob > chi2 = 0.0000
------------------------------------------------------------------------------
dep | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
pre | .1756632 .0533018 3.30 0.001 .0711937 .2801328
treat | -1.759441 .3946169 -4.46 0.000 -2.532876 -.9860058
visit | -.5189469 .0917666 -5.66 0.000 -.6988061 -.3390876
_cons | -.8539732 1.095211 -0.78 0.436 -3.000546 1.2926
------------------------------------------------------------------------------
predict p
table visit treat, cont(mean dep mean p)
------------------------------
| treat
visit | placebo estrogen
----------+-------------------
1 | .8518519 .6176471 /* observed proportion */
| .8917946 .6337553 /* predicted proportion */
|
2 | .8518519 .5588235
| .8336766 .5178159
|
3 | .8148148 .3235294
| .7546193 .4001624
|
4 | .6666667 .3235294
| .6557054 .2923599
|
5 | .5555556 .2352941
| .5431298 .2027277
|
6 | .4074074 .1176471
| .4270517 .1344281
------------------------------
Example with Count Response VariableIn this section we will use data on executions in each of the 50 states for the years 1995, 1997 and 1999.
use http://www.gseis.ucla.edu/courses/data/execute2
describe
Contains data from execute2.dta
obs: 150 2000 us stat abstracts
vars: 7 13 Feb 2002 21:47
size: 4,650 (89.7% of memory free)
-------------------------------------------------------------------------------
storage display value
variable name type format label variable label
-------------------------------------------------------------------------------
sid float %9.0g
state str3 %9s
execute float %9.0g # executions
murder float %9.0g murder rate
unemp float %9.0g unemployment rate
confed float %9.0g confederate state
year float %9.0g
-------------------------------------------------------------------------------
univar sid execute-year
-------------- Quantiles --------------
Variable n Mean S.D. Min .25 Mdn .75 Max
-------------------------------------------------------------------------------
sid 150 25.50 14.48 1.00 13.00 25.50 38.00 50.00
execute 150 1.43 4.74 0.00 0.00 0.00 1.00 37.00
murder 150 5.96 3.31 0.50 3.20 5.85 8.10 17.00
unemp 150 4.67 1.18 2.50 3.70 4.70 5.40 7.90
confed 150 0.22 0.42 0.00 0.00 0.00 0.00 1.00
year 150 1997.00 1.64 1995.00 1995.00 1997.00 1999.00 1999.00
-------------------------------------------------------------------------------
tabstat execute, by(year) stat(n mean var)
Summary for variables: execute
by categories of: year
year | N mean variance
---------+------------------------------
1995 | 50 1.04 8.733061
1997 | 50 1.42 29.14653
1999 | 50 1.82 30.02816
---------+------------------------------
Total | 150 1.426667 22.43418
----------------------------------------
nbvargr execute
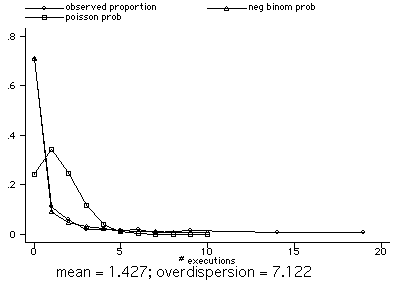 separate execute, by(year)
graph execute1995 execute1997 execute1999 sid, s(iii) c(ll[_]l[-])
separate execute, by(year)
graph execute1995 execute1997 execute1999 sid, s(iii) c(ll[_]l[-])
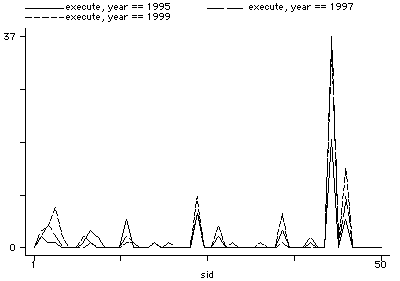 drop murder-confed execute1995-execute1999
reshape wide exec, i(sid) j(year)
(note: j = 1995 1997 1999)
Data long -> wide
-----------------------------------------------------------------------------
Number of obs. 150 -> 50
Number of variables 4 -> 5
j variable (3 values) year -> (dropped)
xij variables:
execute -> execute1995 execute1997 execute1999
-----------------------------------------------------------------------------
corr execute1995 execute1997 execute1999
(obs=50)
| exe~1995 exe~1997 exe~1999
-------------+---------------------------
execute1995 | 1.0000
execute1997 | 0.9481 1.0000
execute1999 | 0.9406 0.9608 1.0000
use http://www.gseis.ucla.edu/courses/data/execute2
xi: nbreg execute murder unemp confed i.year, cluster(sid)
i.year _Iyear_1995-1999 (naturally coded; _Iyear_1995 omitted)
Negative binomial regression Number of obs = 150
Wald chi2(5) = 37.12
Log likelihood = -167.2556 Prob > chi2 = 0.0000
(standard errors adjusted for clustering on sid)
------------------------------------------------------------------------------
| Robust
execute | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
murder | .4056555 .1236221 3.28 0.001 .1633606 .6479504
unemp | -.6013436 .241296 -2.49 0.013 -1.074275 -.1284122
confed | 2.244357 .7560092 2.97 0.003 .7626066 3.726108
_Iyear_1997 | .4385796 .304441 1.44 0.150 -.1581139 1.035273
_Iyear_1999 | .729746 .4136177 1.76 0.078 -.0809298 1.540422
_cons | -1.186817 1.327187 -0.89 0.371 -3.788056 1.414422
-------------+----------------------------------------------------------------
/lnalpha | 1.272271 .2545488 .7733648 1.771178
-------------+----------------------------------------------------------------
alpha | 3.56895 .9084719 2.167046 5.877772
------------------------------------------------------------------------------
test _Iyear_1997 _Iyear_1999
( 1) [execute]_Iyear_1997 = 0.0
( 2) [execute]_Iyear_1999 = 0.0
chi2( 2) = 3.12
Prob > chi2 = 0.2103
xi: xtgee execute murder unemp confed i.year, i(sid) fam(nbin) link(log) corr(exc)
i.year _Iyear_1995-1999 (naturally coded; _Iyear_1995 omitted)
GEE population-averaged model Number of obs = 150
Group variable: sid Number of groups = 50
Link: log Obs per group: min = 3
Family: negative binomial(k=1) avg = 3.0
Correlation: exchangeable max = 3
Wald chi2(5) = 53.39
Scale parameter: 1 Prob > chi2 = 0.0000
------------------------------------------------------------------------------
execute | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
murder | .0860412 .0466163 1.85 0.065 -.005325 .1774075
unemp | -.1054161 .1413111 -0.75 0.456 -.3823807 .1715486
confed | 2.076369 .4320253 4.81 0.000 1.229615 2.923123
_Iyear_1997 | .2408024 .1687292 1.43 0.154 -.0899006 .5715055
_Iyear_1999 | .6461118 .2150903 3.00 0.003 .2245425 1.067681
_cons | -1.040887 .7862023 -1.32 0.186 -2.581815 .5000416
------------------------------------------------------------------------------
test _Iyear_1997 _Iyear_1999
( 1) _Iyear_1997 = 0.0
( 2) _Iyear_1999 = 0.0
chi2( 2) = 9.53
Prob > chi2 = 0.0085
xtcorr
Estimated within-sid correlation matrix R:
c1 c2 c3
r1 1.0000
r2 0.7856 1.0000
r3 0.7856 0.7856 1.0000
drop murder-confed execute1995-execute1999
reshape wide exec, i(sid) j(year)
(note: j = 1995 1997 1999)
Data long -> wide
-----------------------------------------------------------------------------
Number of obs. 150 -> 50
Number of variables 4 -> 5
j variable (3 values) year -> (dropped)
xij variables:
execute -> execute1995 execute1997 execute1999
-----------------------------------------------------------------------------
corr execute1995 execute1997 execute1999
(obs=50)
| exe~1995 exe~1997 exe~1999
-------------+---------------------------
execute1995 | 1.0000
execute1997 | 0.9481 1.0000
execute1999 | 0.9406 0.9608 1.0000
use http://www.gseis.ucla.edu/courses/data/execute2
xi: nbreg execute murder unemp confed i.year, cluster(sid)
i.year _Iyear_1995-1999 (naturally coded; _Iyear_1995 omitted)
Negative binomial regression Number of obs = 150
Wald chi2(5) = 37.12
Log likelihood = -167.2556 Prob > chi2 = 0.0000
(standard errors adjusted for clustering on sid)
------------------------------------------------------------------------------
| Robust
execute | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
murder | .4056555 .1236221 3.28 0.001 .1633606 .6479504
unemp | -.6013436 .241296 -2.49 0.013 -1.074275 -.1284122
confed | 2.244357 .7560092 2.97 0.003 .7626066 3.726108
_Iyear_1997 | .4385796 .304441 1.44 0.150 -.1581139 1.035273
_Iyear_1999 | .729746 .4136177 1.76 0.078 -.0809298 1.540422
_cons | -1.186817 1.327187 -0.89 0.371 -3.788056 1.414422
-------------+----------------------------------------------------------------
/lnalpha | 1.272271 .2545488 .7733648 1.771178
-------------+----------------------------------------------------------------
alpha | 3.56895 .9084719 2.167046 5.877772
------------------------------------------------------------------------------
test _Iyear_1997 _Iyear_1999
( 1) [execute]_Iyear_1997 = 0.0
( 2) [execute]_Iyear_1999 = 0.0
chi2( 2) = 3.12
Prob > chi2 = 0.2103
xi: xtgee execute murder unemp confed i.year, i(sid) fam(nbin) link(log) corr(exc)
i.year _Iyear_1995-1999 (naturally coded; _Iyear_1995 omitted)
GEE population-averaged model Number of obs = 150
Group variable: sid Number of groups = 50
Link: log Obs per group: min = 3
Family: negative binomial(k=1) avg = 3.0
Correlation: exchangeable max = 3
Wald chi2(5) = 53.39
Scale parameter: 1 Prob > chi2 = 0.0000
------------------------------------------------------------------------------
execute | Coef. Std. Err. z P>|z| [95% Conf. Interval]
-------------+----------------------------------------------------------------
murder | .0860412 .0466163 1.85 0.065 -.005325 .1774075
unemp | -.1054161 .1413111 -0.75 0.456 -.3823807 .1715486
confed | 2.076369 .4320253 4.81 0.000 1.229615 2.923123
_Iyear_1997 | .2408024 .1687292 1.43 0.154 -.0899006 .5715055
_Iyear_1999 | .6461118 .2150903 3.00 0.003 .2245425 1.067681
_cons | -1.040887 .7862023 -1.32 0.186 -2.581815 .5000416
------------------------------------------------------------------------------
test _Iyear_1997 _Iyear_1999
( 1) _Iyear_1997 = 0.0
( 2) _Iyear_1999 = 0.0
chi2( 2) = 9.53
Prob > chi2 = 0.0085
xtcorr
Estimated within-sid correlation matrix R:
c1 c2 c3
r1 1.0000
r2 0.7856 1.0000
r3 0.7856 0.7856 1.0000
Categorical Data Analysis Course
Phil Ender